Cancer research has gone through a process of development in the last thirty years due to the growth in the field of genetics. Before the advent of genomic testing, most individuals with a certain type and stage of cancer received the same chemotherapy, but these treatments had variable outcomes. The improvement in genetic technologies; however, allowed clinicians to find dissimilarities in patients’ tumors of the same cancer type, and identified different signalling pathways to target therapeutically. Such observations underpinned personalised medicine as well as aided personalised cancer risk assessments.
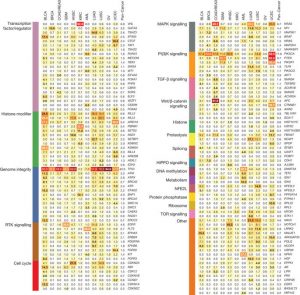
Mutational landscape across 12 major cancer types. Kandoth et al. (2013). Nature, 502(7471):333-339.
Marty et al investigated the latter from a completely different perspective. Instead of focusing on driver mutations and genes previously implicated in oncogenesis (‘cancer genes’), they studied the major histocompatibility complex (MHC) that has a crucial role to prevent tumor progression. MHC-I molecules are expressed on all nucleated cells and through displaying intracellular protein content on the cell surface, they allow the immune system to detect oncogenic changes and eliminate neoplastic cells. Of note, MHC-I molecules are highly polymorphic and have different binding affinities for different peptides. The authors found that the MHC-I alleles someone carries determine which oncogenic alterations are more likely to escape immune surveillance as a result of their poor binding. It means that individuals who have higher risk for certain types of cancer could be identified based on their MHC-I genotypes.
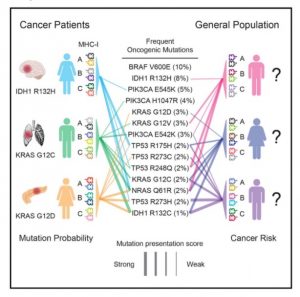
MHC-I genotype is associated with the appearance of specific oncogenic mutations, and recurrent oncogenic mutations are biased toward poorly presented peptides. Marty et al. (2017). Cell, 171(6):1272-1283.
This pioneering study on the major histocompatibility complex from the perspective of cancer susceptibility will likely catalyze further investigations. Once databases will be available on the correlation between HLA-1 genotypes and their binding affinities for known oncogenic peptides, they will suggest routine HLA genotyping for personalised disease prevention. Even more so, these findings might explain why certain individuals with strong genetic risk factors for cancer do not develop the disease phenotype – owing to their HLA architecture that has a better binding affinity for the altered protein. Hence it warrants for a combined screening of driver mutations and HLA genes to get a closer estimation of disease risk.
Most importantly though, the findings of Marty et al emphasise the clinical utility of high resolution HLA screening way beyond the field of transplantation.
Dorottya M Berki, PhD (dorottya.berki@omixon.com) Internal Support Scientist at Omixon
References:
Marty et al. (2017). MHC-I Genotype Restricts the Oncogenic Mutational Landscape. Cell 171: 1–12.
Kandoth et al. (2013). Mutational landscape and significance across 12 major cancer types. Nature, 502(7471):333-339.
