The following interview is part of our interview series that we are conducting with Key Opinion Leaders, Experts and Business Managers within the HLA field. This time, Carol Kneibb and Ryan Morlen from Children’s Hospital of Philadelphia talk about their experiences and thoughts on HLA laboratory activities. We hope this series will shed light on trends and challenges within the profession and will be helpful to all HLA experts.
Carol Kneibb (C.K.) and Ryan Morlen (R.M.) have had very different paths prior to their employment in the field of Immunogenetics. Ryan has moved from various labs as an established laboratory specialist whereas Carol has spent the last 22 years of her life devoted to HLA in the high-volume labs in Curitiba, Brazil. Now Carol and Ryan both work at the Children’s Hospital of Philadelphia under Dr. Dimitri Monos, where she supervises the SOT program and Ryan is a Technical Specialist. Carol has developed a special talent that many HLA labs are just now beginning to uncover its significance in the prediction of organ rejection. Her skill for epitope analysis has been fine-tuned throughout her extensive career and upon bringing Carol’s expertise to CHOP’s lab, where all samples are typed via NGS, Carol now has the data she always dreamed about when in Brazil. Combining her skills with the abundant CHOP data, Ryan’s ingenuity, and guidance from Dr. Monos, Carol has created some very impressive tools that are currently being clinically implemented in CHOP’s lab. These Virtual Crossmatch (VXM) tools can give clinicians more data and are a huge leap towards a faster, more accurate, and truly personalized VXM.
What is your background and how did you come to HLA?
C.K. – I have a BSc in biology with a M.Sc in Health Sciences. I heard about HLA for the very first time during my sophomore year in college when I found out a friend of a friend was working in a lab called Immunogenetics. This word called my attention and I started to search about it. Finally, in my senior year I was able to have classes about it and I got an internship in the same Immunogenetics Lab I had heard about before. It was in 1999!
R.M.– My background has been quite convoluted. My original field of study in college was computer science, however, after my mom became ill, I relocated to be closer to my family. After relocating, my interest started to move towards biological sciences and I started studying microbiology and laboratory medical sciences. Throughout my education, I was never exposed to HLA. After my wife completed her medical school training, we relocated from Washington to Delaware and I came across an open position at the Children’s Hospital of Philadelphia in the Immunogenetics Laboratory. For myself, learning HLA and being introduced to HLA was somewhat happenstance.
What is it that you enjoy about HLA?
C.K. – HLA became my passion. It was something I learnt how to love. My main interest is the serology part of it. The immunology behind the entire system. You can have a standard pattern of reaction, but every person, every transplant can respond in a different way. I love the challenges we have with the cases where the information in the books is not applicable. Every case is basically unique.

Ryan Morlen, CHOP
R.M. – The HLA community is what I enjoy the most about HLA. The community is very tight-knit and the laboratory makes dramatic impacts on the well-being of patients that are awaiting or have received transplants. Additionally, I’ve gotten to lead/work on some very cool projects ranging from automating the wet-bench setup of our NGS assay to developing a web tool for adding additional analytical depth to our solid organ transplantation workflow.
Epitope Analysis is a heavily argued topic in terms of how to analyze and implement the data clinically. Where do you stand in this debate?
C.K. – I started to work with epitopes in 2003 with the first version of triplets. In my old lab in Brazil, we used it for antibody reactivity analysis. At that point, we were using ELISA as our methodology. Since then, a lot of things have changed. We started using LABScreen and for each new methodology we adapted our epitope analysis. All the PRA tests in the lab were analyzed with epitope analysis as well. At that point, we had the cut off of 500 MFI, however, each patient had his/her own cut off determined according to the antibody reactivity pattern against epitopes. As the years passed, we were able to work together with other labs and they developed a web tool that helps to perform the epitope analysis. At this point we were using the new version of eplets, combined with epitopes from Terasaki and CREG groups. Basically, since 2003, I never spent one day without working with epitopes. Today, I have the opportunity to work with high resolution typing by NGS, which makes my life as an epitope analyzer easier, and gives me the chance to add the amino acid analysis to my work as well.
Based on everything I learnt through the years and analysis I did, today, with more tools and a little bit more knowledge, I think you need to understand better the concept of epitopes to know where and when to use it. It can be “the” help for challenging cases, but it can also be irrelevant in other cases. I am definitely not using it for every case as I did in the past, but I still use it every day. In my experience, I think we should use epitope analysis as a tool to personalize the transplant immunology analysis.
R.M. – I personally believe that epitope analysis will be or should be very important in the near future. While epitope analysis is not yet a perfect science, the underlying principles when applied correctly could profoundly impact the way organs are allocated in the future. Some of this is already seen across European transplant sites where epitope matching is used to expand the limited donor pool. At worst, we should consider epitope analysis as another tool at our disposal for providing the best possible care to transplant patients.
How can you use the data acquired from epitope analysis?

Carol Kneibb, CHOP
C.K. – Basically we use it in 3 scenarios. First, to determine on a deeper level the compatibility between patient and donors, and access permissible mismatches. Second, to analyze the antibody reactivity pattern against an epitope(s) and not only against a single antigen. And third, to determine surrogate beads in order to monitor donor specific antibodies when the donor’s specificity is not available in the PRA panel in use.
R.M. – The data from epitope analysis can be applied in several ways. We can look at similarities and differences between patients and donors, similarities and differences between alleles, and begin to explain reactivity patterns from Luminex Single Antigen Beads (similar to CREGS). More interesting perhaps might be the ability to look beyond the limited alleles present within the Single Antigen Bead panels and perform what could be referred to as a surrogate bead analysis. In a surrogate bead analysis, we substitute a bead in the Single Antigen Bead panel for a donor allele which is not available but can be considered identical at the molecular level (considering the differences between patient and donor).
How does epitope data further personalize/enhance the VXM?
C.K. – When you use epitope analysis in the antibody reactivity scenario, you can increase your chance to predict a positive crossmatch. This can happen because between all antibodies against that specific epitope, you will have your DSA and others, and since the reactivity is against an epitope, you can use the highest MFI value as DSA, even though the antigen isn’t from the donor.
R.M. – Over time there has been a steady evolution in the way matching has occurred in transplantation. First, we progressed from antigen to allele — now a methodology exists which can take us one step further. When we identify reactivity against epitopes, we often can include or exclude entire groups of alleles at one time. From a VXM standpoint, we’re able to make the best possible bead selections for a patient based on their entire history rather than a single instance. Making the best choices early, say in a VXM situation, allows the best patient to receive an organ and hopefully prolong the life of their transplant.
What are your goals with the VXM?
C.K. – For the patient: to minimize unnecessary testing, and patient’s and donor’s blood collections. For the lab: to make this tool even more effective and useful. VXM is a great source of analysis for a patient’s immunologic history against the donor’s antigens.
R.M. – My goals for the VXM are to make it as simple and accurate as possible. There is no reason that a laboratorian should ever need to flip through paper charts or compile spreadsheets at 3 am in order to provide a VXM. We’ve worked hard to build tools to automate the process and provide confidence in the underlying data to which these critical transplant decisions are being made. Laboratories should truly reflect on their processes and attempt to minimize manual data manipulation wherever possible as a safety initiative not only for patients, but staff as well.
Where do you see the VXM in the next 5 years?
C.K. – I would like to see it as the primary test for patient versus donor. A physical XM would be performed according to VXM results, if needed.
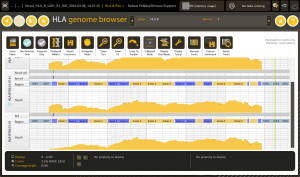
HLA genome browser
R.M. – The VXM will be with us for a long time to come. VXMing is a very quick and easy analysis when the answer is a clear yes or no but tends to be ambiguous for the in-between cases. It would be nice to see an industry and community-driven initiative to consolidate and build tools for standardizing and performing VXMs. Each institution is currently left to develop and implement its own workflow which may vary dramatically from location to location. It would be advantageous for the HLA community as a whole to address our deficiencies and qualifications when it comes to VXMs so we may provide not only the best care for transplant patients but also provide rapid safe results for clinicians who are making these decisions.
Do you think we will reach a point in the near future where we can forgo FXM due to such thorough VXMs?
C.K. – No. We still have some issues with the antibody screening testing in which we can have false positive reactions and denatured antigens, where a physical XM (with the real donor or even a surrogate donor) is very important.
R.M. – For the surgeons and other physicians that will be performing the transplant and managing the care of a patient, the prospect of replacing a physical crossmatch with a virtual one is likely intimidating. While feasible from a technical standpoint, performing a virtual crossmatch in lieu of a physical crossmatch is about risk management. Clinicians would need to establish new guidelines and study outcomes of patients from multiple cohorts before replacing a physical crossmatch with a virtual one. It may not be unsafe to treat positive VXMs as positive FXMs, but the question remains how can we help patients for which the VXM is simply incorrect or have rejection due to non-HLA factors? Those are the primary hurdles that must be overcome before we can seriously tackle this question.
How do you use the VXM now that differs from other labs?
C.K. – We are trying to implement (with success) the VXM as a primary test for kidney living donors. The patient and donors are first HLA typed by NGS and a VXM as well as mismatch analysis is performed to help the transplant program find the better donor.
R.M. – What I believe differentiates our laboratory from others concerning performing VXMs is our workflow. We’ve created a semi-automated process that eliminates transcription errors and allows us to perform VXMs very quickly. Once we resolve the donor’s two-field typing (using the NMDP’s HaploStats and antigen level typing from UNOS), a technologist only needs to select the appropriate beads to match the donor’s type. Our solution communicates directly with our HistoTrac database and automatically sorts the patient’s Single Antigen Bead data into the current and highest historical values after simply selecting the specificity. The data is presented in both a tabular and graphical manner that gives a complete view of the patient’s potential risk to the clinician across all serum samples the laboratory has ever processed.
The process has been further automated where all specificities are automatically chosen and beads that are missing from the Single Antigen panel are chosen based on their similarity at the amino acid level to the patient and donor mismatches. We are looking forward to implementing the new process in the near future.
HLA Twin
Why is NGS crucial for accurate epitope analysis?
C.K. – The HLA typing resolution is very important for epitope analysis. HLA typing by NGS increases the level of the epitope analysis accuracy. There are a lot of cases where the most frequent allele, determined by haplotype or allele frequencies for low/medium resolution typing, is not the correct allele. In this case, this difference can interfere in the epitope analysis. Furthermore, since not all the epitopes have their immunogenicity known, the complete information of HLA typing can help to guarantee the correct epitope determination.
R.M. – As Next Generation Sequencing becomes more and more prevalent in HLA laboratories, so will more advanced solid organ transplantation analysis methods. When considering epitope analysis, knowing which alleles are involved is critical because many alleles comprise say, the A2 antigen. The differences between A*02:01 and A*02:02 are really what we are looking at when we start to consider epitopes. Without Next Generation Sequencing data or other high-resolution typing data, we are left to make assumptions about which alleles we are looking at. Having high-resolution typing, however, allows us to make the most accurate predictions possible.
We would like to say thank you to Carol and Ryan for offering their time and answered our questions!
Previous interview:

